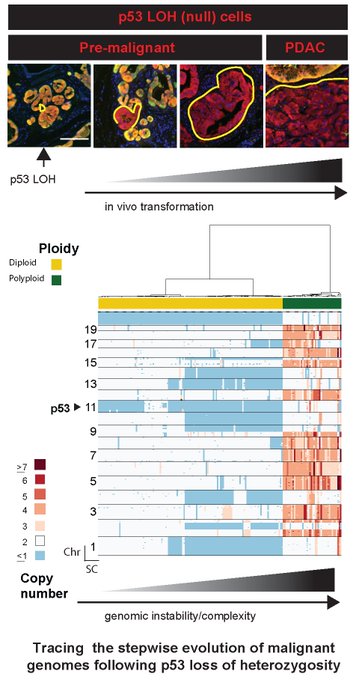
Cells that lose p53 undergo a deterministic pattern of genome evolution with the following order: p53 Loss Of Heterozygosity -> accumulation of Chromosomal Deletions -> Genome Doubling -> Gains & Amplifications. 4/n
1
4
27
Replies
Out in
@Nature
today, we report that
#p53
loss unleashes an ordered & selective pattern of cancer
#GenomeEvolution
: . 1/n
11
242
936
30 yrs ago,
#p53
was dubbed ‘Guardian of the Genome’ based on ability to prevent
#GenomeInstability
and, indeed, p53-mutant tumors are often heterogenous, genomically rearranged & polyploid. But how do these genomes evolve? 2/n
1
0
12
To answer this question, we leverage a unique
#MouseModel
that enables tracing of cells that lose p53 as they cross the benign-to-malignant transition. p53 loss is NOT enough to promote cancer; instead, other events associated with further genome evolution are required. 3/n
1
3
16
Our results show how ‘genome instability’ following loss of the Guardian contributes to
#TumorEvolution
and the acquisition of malignant features; they also have implications for the aggressive nature of
#p53
-mutant tumors and strategies to target them. 5/n
1
1
12
Congratulations to lead authors
@TimourBaslan
,
@JohnPMorrisIV
& Zhen Zhao
@sparklingpm
, and many thanks to all of our spectacular collaborators
@LabNotta
,
@Nevenka_D
,
#SteveLeach
@DartmouthCancer
,
@ciacobu
@CpcrMSK
,
@MskccCMO
@MSKCancerCenter
! 6/n
1
3
25